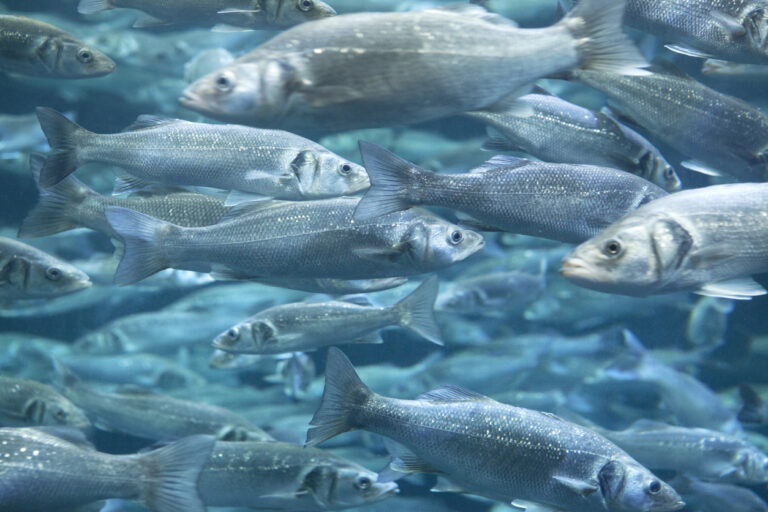
Identifying and analyzing biomarkers in animal health
For many years, DIAG4ZOO has been identifying and analyzing biomarkers in animal health, and developing innovative diagnostics in partnership with players in the marine industry.
These diagnostics are intended for routine use in aquaculture in order to improve animal health monitoring, farm quality and optimize production: early detection of pathogens, selection of individuals of interest and resistant progenitors, targeted use of antibiotics, evaluation and control of culture media in closed or open circuits (microbiomes) and sanitary monitoring of these circuits. These tests help to optimize biosecurity approaches for aquaculture production: fishes, seashells, crustaceans, etc.
These diagnostics are based on the upstream identification of DNA or RNA biomarkers, and can be used on existing automated systems (PCR, real-time PCR), or directly in the field (strip test).
The scientific team’s expertise in biomarker identification is not dependent on animal species. Thus, its know-how in test design (PCR or field) is applicable to any marine species, and for a wide range of applications.
The Diag4Zoo team has worked in animal health on fishes, oysters, shrimps and certain molluscs, in the fields of food safety (health monitoring), agri-food, environment (assessing the impact of pesticides or herbicides in the marine environment), and in the development of new active compounds.
These studies are generally the fruit of collaborations with academic and private teams, notably with partners in Occitanie: Ifremer, CIRAD, Cepralmar, CRCM (Mediterranean Regional Shellfish Growing Committee), BARBA group…
Fishes : sea bass / sea bream / trout / tuna/ swordfish



Virus detection
The Diag4zoo team has developed a detection test for Nodavirus, a virus that can affect various types of farmed fishes: sea bass, sea bream, grouper…
The detection kit or analysis service provided is based on enzymatic amplification of a specific nodavirus sequence by PCR. This technology is highly sensitive and delivers results in less than 4 hours.
Biomarker identification
The Diag4zoo team has also carried out studies on trout, identifying biomarkers correlated with the animal’s diet and environment.
Species identification
DIAG4ZOO has developed an innovative method for identifying tuna species from meat samples (loin, steak…), using a SNP-rich region of the ND4 mitochondrial DNA (mtDNA) gene.
This genotyping approach (PCR amplification + sequencing) allows to obtain a very rapid result and enable the precise differentiation of economically important and different species such as Yellowfin Tuna (T. albacares), Bigeye Tuna (T. obesus) or Atlantic Bluefin Tuna (T. Thynnus).
This tuna species identification service is now available to industry professionals.
Using genomic databases and its tools, the company’s scientists have identified two specific regions of the swordfish genome, enabling them to design a technique for the genetic identification of swordfish to ensure its traceability. The methodology used is based on the principle of DNA Barcoding (developed by Hebert et al., 2009), which is currently the reference method for species identification.
In general, the use of genetic traceability methods to identify certain species can be essential for food and distribution professionals, enabling them to carry out quality controls on purchased products.
Thanks to DIAG4ZOO’s technical skills and know-how in the rapid development of new genetic traceability methods, professionals benefit from technical expertise applied to veterinary products, to validate their labelling and ensure the traceability of products delivered by their suppliers.
Mediterranean oyster
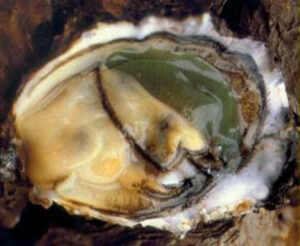
Diag4zoo has collaborated with Ifremer (Montpellier), Cepralmar (center for the study and promotion of lagoon and maritime activities), and CRCM (Comité Régional Conchylicole de Méditerranée) on the identification and selection of Mediterranean oysters resistant to pathogens and summer mortalities.
The studies focused on the molecular mechanisms and genetic bases involved in the survival of Crassostrea gigas oysters to infections by virulent Vibrio, such as V. splendidus LGP32 and V. aestuarianus 02/41 Lpi. These infections have been observed frequently in farmed oysters and have led to significant mortalities in Mediterranean oyster production.
Thanks to high-throughput genomics and transcriptomics technologies, and based on reproducible and standardized laboratory experiments, the analyses led to the identification of a combination of biological markers that could predict the ability of oysters to survive to Vibrio infections.
This combination of biological markers, established in a first stage on the basis of laboratory experiments, was then validated in a second stage on the basis of analyses carried out on oysters present in the natural environment. This combination of biological markers enabled us to distinguish between oysters capable of surviving episodes of major mortality in the Thau lagoon and those that will die, with 73% reliability.
In addition, a second study was carried out to investigate the ability of oysters to transmit these survival markers to their descendants, genetic traits conferring better survival capacities to Vibrio infections.
This selection approach, based on the identification of biological markers, could be applied to produce oyster “lines” adapted to different environmental conditions and/or resistant to different pathogens. In this way, the production of local strains adapted to the specific environmental conditions of different oyster farming areas could help reduce the risks of disease and stress in oysters.
Against a backdrop of mortalities leading to significant economic losses, professionals could produce oysters resistant to certain infections and adapted to certain environmental conditions. Ultimately, these new practices would make the shellfish industry more secure.
Pearl oyster from French Polynesia
Pearl farming is an important French aquaculture activity. It is the French Polynesia’s second largest export resource, after tourism.
Diag4zoo collaborated with Ifremer (Tahiti) and various stakeholders involved in pearl farming to conduct researches aimed at improving pearl quality.
The studies focused on
- the selection of high-performance pearl oyster lines, which involves controlling reproduction and cross-breeding,
- understanding the genetic determinism of pearl color and the mechanisms leading to the appearance of pearl surface defects.
The evolutionary success of molluscs can be attributed in part to their efficiency in supporting and protecting their soft bodies with a biomineralized external structure, the shell. However, the knowledge of the proteins responsible for the shell’s microstructural polymorphism formation and its unique properties is still very fragmentary.
The Diag4zoo scientific team has therefore been working on the different secretory repertoires that control the processes of biomineralization of the prism and nacre deposition of the pearl oyster shell.
In Pinctada margaritifera and Pinctada maxima oysters, 80 shell matrix proteins have been identified, 66 of which are entirely unique. This is the only description of the entire matrix “biomineralization toolbox” that is thought to regulate the formation of prismatic and pearly shell layers in pearl oysters. These studies have demonstrated that prisms and nacre are assembled from very different protein repertoires, suggesting that these layers do not derive from each other.
The shell of the pearl oyster Pinctada margaritifera is composed of an organic matrix that plays a key role in the dynamic process of biologically controlled biomineralization. To better understand this process, transcriptome and proteome analyses of the calcifying mantle and shell of Pinctada margaritifera were carried out.
To expand the catalog of genomic data and identify shell matrix proteins involved in biomineralization in P. margaritifera, certain fractions of the oyster genome were sequenced (Expressed Sequence Tag, EST) corresponding to calcifying mantle tissue. These data were then combined with a proteomic analysis of the shell.
These analyses made it possible to cover most of the diversity of P. margaritifera shell matrix proteins, but also showed that oyster mantle transcripts code for proteins present in the shell, demonstrating their involvement in shell formation.
The combination of transcriptomic and proteomic approaches is therefore a powerful means of identifying the proteins involved in biomineralization. The data generated in this study provided a comprehensive list of sequences linked to biomineralization, and enabled a major breakthrough in the field of mollusc biomineralization.
Shrimp
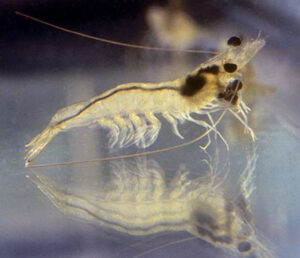
Biomarker identification
The Diag4Zoo team collaborated with Ifremer (Montpellier) to identify biological markers linked to the ability of shrimp to survive to vibrio infections.
Understanding the antimicrobial defense mechanisms of shrimp would thus help in the design of effective strategies for disease management and control in aquaculture.
In this study, transcriptomic analyses of circulating antimicrobial peptide (AMP) hemocytes were carried out. A relationship was found between the expression level of certain antimicrobial peptides and the positive response of Litopenaeus stylirostris shrimp to resist pathogenic infection by Vibrio penaeicida. In particular, significant differences in certain antimicrobial peptides were demonstrated between unselected shrimp and the third generation of a shrimp line selected for their ability to survive to Vibrio penaeicida infections.
Microbiome / Microbiota
The Biofloc technology (BFT) is a farming method requiring little or no water exchange. It is widely used in aquaculture, particularly in shrimp production. In the water column, these systems develop conglomerates of microbes, algae and protozoa, as well as detritus and dead organic particles. The microbial community present in these systems can be used as a pond water treatment system, and the microbial proteins can be used as a food additive.
The current problem with Biofloc technology is the difficulty of controlling the composition of its bacterial community to obtain optimum water quality and guarantee shrimp health.
The aim of the study, in which the Diag4Zoo scientific team took part, was to use sequencing in order to study the microbial diversity (or microbiome) present, on the one hand, in water samples from two culture environments (Biofloc and seawater) and, on the other, in the intestines of shrimps raised in both environments.
Analyses of the bacterial community identified in waters from the two environments, BFT system and “clear seawater”, containing the shrimp Litopenaeus stylirostris, revealed large differences in the frequency distribution of operational taxonomic units (OTUs). Four of the five most dominant bacterial communities were different in the two culture environments. Bacteria found in high abundance in BFTs have two main characteristics: the need for an organic substrate or nitrogen sources to grow, and the ability to attach to surfaces and aggregate. A correlation was found between bacterial groups and physicochemical and biological parameters measured in the rearing ponds. In addition, the bacterial communities in the rearing water influenced the shrimp microbiota. Indeed, the Biofloc environment modified the shrimp’s intestinal microbiota, as shown by the low level (27%) of similarity between intestinal bacterial communities in the two environments.
This study provided the first information describing the complex microbial community of the Biofloc system, and could help to understand the environment-microbiota-host relationship in this farming system.
These interactions are being applied to research into immunity, disease resistance and nutrition in farmed shrimp.
The Diag4Zoo team also uses sequencing and specific bioinformatics treatments to identify and monitor specific markers of the gut microbiota in relation to the immune response or resistance of animals to certain pathologies or infections.
Two technical approaches are used to analyze the intestinal microbiota:
- 16S ribosomal RNA gene sequencing,
- analysis of all bacterial genes using quantitative metagenomics.
Molluscs: sea cucumber, sea cone
Sea cucumber
The Diag4zoo team regularly carries out analyses to detect the presence of Nodavirus in marine animals, in particular sea cucumbers “Holothuria tubulosa”. The virus is detected by qPCR on gonad samples: the technology used is highly sensitive, enabling a response and a result to be obtained in less than 4 hours.
Sea cone
The marine cone is a type of sea snail, a predatory gastropod mollusk that possesses one of the most fearsome venoms in the marine world. This sea cone venom contains a complex blend of bioactive peptides to subdue the animal’s preys.
The DIAG4ZOO scientific team took part in an international study using a combination of proteomic and transcriptomic approaches to discover a set of conopeptides and conoproteins present in the injectable venom of Conus consors, a fish-hunting Indo-Pacific cone snail.
From a large set of venom gland transcriptomic mRNA sequences identified in the course of this study, 105 components were isolated out of some 400 molecular masses detected in the venom. Among these components, new conotoxins were identified and described, as well as a new family of disulfide-free conopeptides. These results provide the most comprehensive and accurate proteomic overview to date of an injectable cone snail venom, and delineate the main protein families present in administered venom.
This study demonstrates the feasibility of this analytical approach, and paves the way for strategies to discover new therapeutic molecules through the use of transcriptomics in animals.