DIAG4ZOO has positioned itself as an expert in the field of microbiome analysis of samples from aquaculture production.
In a context of climate change, food safety and population monitoring, microbiome analysis in aquaculture plays a major role in preventing disease or mortality. Moreover, the impact of the microbiome extends far beyond this, as it plays a decisive role in fish quality of life. Indeed, the health of the microbiome directly influences fish growth and flesh quality, crucial aspects in the aquaculture production sector.
DIAG4ZOO was commissioned by a major European aquaculture producer to study the ecosystem of its ponds, i.e. to analyze all the microorganisms (bacteria, viruses, fungi, yeasts) predominating in the environment under study.
To carry out these microbiome analyses, DIAG4ZOO took an integrative approach of the viral and bacterial communities (microbiomes) present in different samples, such as eggs, larvae and fish (gills and fins via non-lethal sampling). In addition, the animal’s aquatic environment, i.e. the water, is also meticulously analyzed.
Microbiome analysis enable to identify and count micro-organisms by isolating the genomes present in samples (water, larvae, feces, etc.), sequencing them and identifying which organisms these genomes are associated with.
Microbiome analysis method
The metagenomic analysis process followed by DIAG4ZOO involves several methodical steps, ensuring maximum accuracy and completeness. The method consists of the following successive phases:
- Extraction and purification of the DNAs : This step aims to efficiently isolate the genetic material required for analysis.
- Qualification of the DNAs : DNA quality assessment eliminates any undesirable elements and ensures the reliability and reproducibility of the genetic data to be analyzed.
- PCR amplification and sequencing : qualified DNAs are amplified by polymerase chain reaction (PCR), then subjected to high-throughput sequencing on a state-of-the-art platform. This step enables to obtain a considerable amount of genetic data for analysis.
- Bioinformatics and biostatistical analysis of the sequence data : the sequence data obtained is subjected to rigorous analysis, combining advanced bioinformatics techniques and proprietary biostatistical methods. This makes it possible to decipher the genetic information and extract meaningful conclusions..
DIAG4ZOO adopts a targeted approach, depending on the expected objectives and their purpose (spot check or implementation of a routine monitoring tool). Its technical team may choose to focus on a variable region of the 16S ribosomal RNA gene, or to analyze the entire ribosomal DNA.
In the present study, the first approach (16S ribosomal RNA gene sequencing) was preferred. Targeted metagenomic sequences from the microbiome were analyzed using the bioinformatics pipeline developed by DIAG4ZOO.
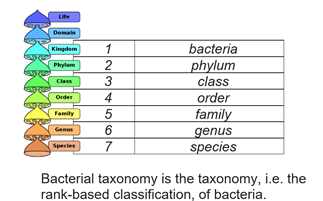
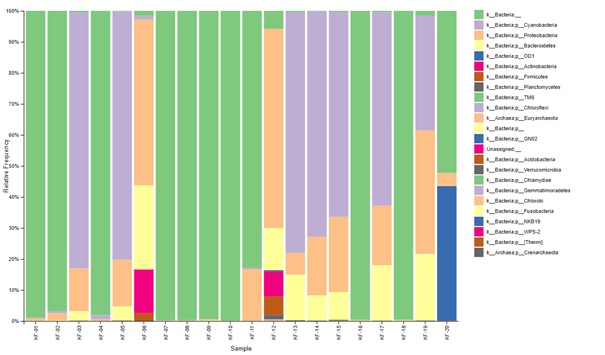
This work led to the identification of the structuring of prokaryotic communities, also known as operational taxonomic units (OTUs), within the various samples.
The end result offers a classification of the readings at several taxonomic levels, encompassing phylum, class, order, family, genus and species, thus providing an in-depth understanding of the microbiological diversity present in the samples studied.
Thus, DIAG4ZOO, through its in-depth expertise and integrative approach to microbiome analysis, is making a significant contribution to improving the health and productivity of farmed fish, responding to contemporary challenges linked to sustainability, food quality and the rational management of aquatic resources.
More generally, DIAG4ZOO’s scientific team has considerable expertise and experience in the Barcoding and Metabarcoding approaches, two sophisticated techniques based on the extraction and high-throughput sequencing (NGS) of DNA.
In this respect, the DIAG4ZOO’s team excels in the art of meticulously extracting DNA from various samples, whether homogeneous or heterogeneous. By then using high-throughput sequencing technologies, the team is able to obtain a significant amount of genetic data quickly and accurately.
These Barcoding and Metabarcoding approaches are proving particularly powerful for the analysis of a variety of samples, even heterogeneous ones, paving the way for multiple applications:
- Rapid and specific detection of pathogens present in samples, to reinforce disease prevention and management actions;
- Monitoring of Invasive Species, providing crucial information for ecosystem preservation;
- Detailed monitoring of Taxonomic Groups, providing a better understanding of biological diversity within samples;
- Diet studies, providing valuable information on ecological interactions and food chains.
DIAG4ZOO’s expertise in Barcoding and Metabarcoding techniques can thus contribute significantly to the advancement of knowledge in crucial fields such as environmental health, biodiversity preservation and food safety.
To find out more about DIAG4ZOO’s expertise in microbiome analysis, metagenomic analysis, Barcoding and Metabarcoding approaches, genome or DNA sequencing, in aquaculture or in complex samples, please contact our sales team.
